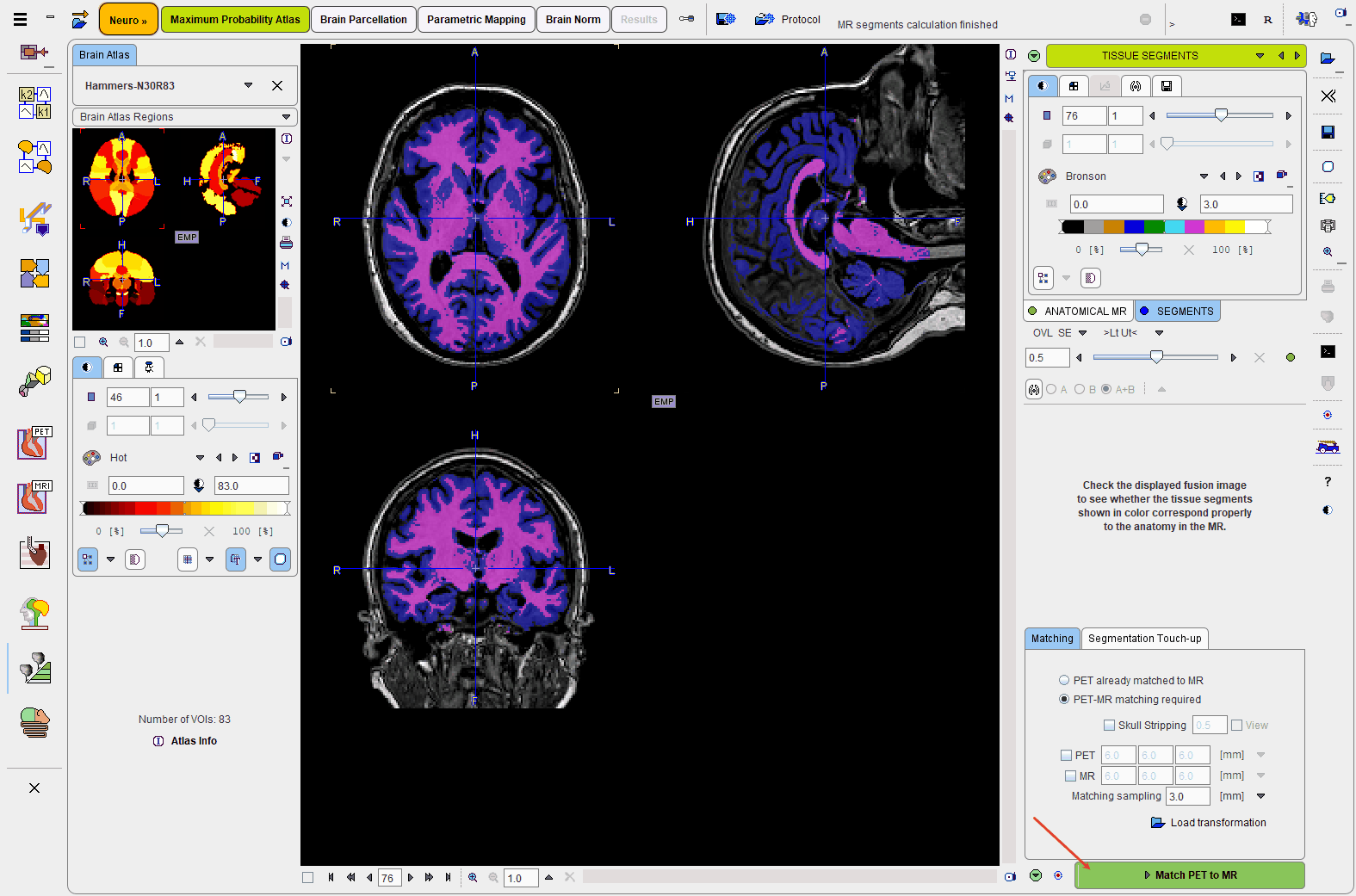
The result of the segmentation is shown as a fusion of the tissue segment map with the MR image on the TISSUE SEGMENTS page. Note that the TISSUE SEGMENTS image tab contains a label image with gray matter, white matter and CSF represented by the label values 1, 2 and 3, respectively.
Segmentation Touch-up
The label map is calculated from the probability maps of GM, WM and CSF. The relative extent of the tissue categories can be modified on the Segmentation Touch-up panel using two different methods. Any modification is immediately reflected in the display.
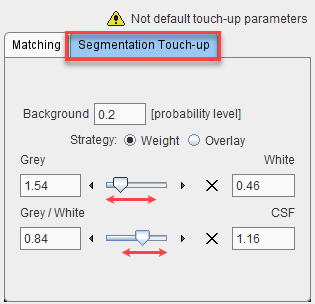
If the probability value in all of the GM, WM, and CSF maps is below the Background level, a pixel is classified as background and assigned the background label value of 0.
With the Weight strategy the boundary between GM and WM can be shifted. The following procedure is applied in all non-background pixels: The GM and WM probabilities are multiplied by the factor in the Grey/White field, and the CSF probability by the value in the CSF field. If the scaled probability of CSF is higher than the scaled GM and WM probabilities, the pixel is assigned the CSF label value of 3. Otherwise, a similar comparison is done between GM and WM: The GM probability is multiplied by the factor in the Grey field, and the WM probability by the value in the White field. If the scaled GM probability is higher than the scaled WM probability, the pixel is assigned the GM label value of 1, otherwise the WM label value of 2.
The Overlay strategy simply uses two thresholds for the GM and the WM probability map, with white matter having higher priority.
Please use the fusion slider to evaluate the segmentation quality. In case the result is not satisfactory, return to the previous page, modify the segmentation parameters, then activate Segment T1 MR again.
PET to MR Matching
The next step consists of rigidly matching the averaged PET image to the MR image, with the options
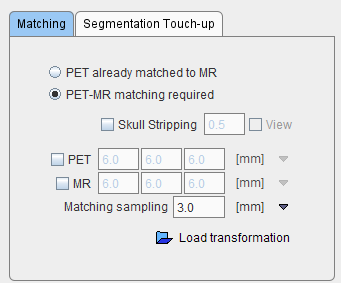
If the data is already matched, the calculation can be skipped by activating the PET already matched to MR radio button. If the matching has been performed before and the transformation saved, it can be loaded and applied with the Load transformation button.
Otherwise, PNEURO will apply a rigid matching procedure based on the Normalized Mutual Information criterion with Matching sampling as the main parameter. Optionally, if the result is not satisfactory, the PET and/or the MR image may be smoothed.
Please activate the Match PET to MR action button to start matching.