The 2-Tissue (BFM) model implements fitting a two-tissue compartment model in each image pixel. It is based on an analytic solution of the system of differential equations which results in the calculation of two eigenvalues α1 and α2.
![]()
The expected tissue activity is obtained by the convolution of the input function with a sum of two decaying exponentials plus a contribution from whole blood.
![]()
For FDG, irreversible binding can be assumed and k4 set to zero, whereby α1 becomes zero. Hereby the number of fitted parameters is reduced and the operational equation simplifies to
![]()
Overview of the BFM Processing in PXMOD
In the MRGlu (FDG BFM, Slice-dependent Times) model, sonly the irreversible configuration is supported. It uses the simplified basis function solution with fits 4 parameters: θ1, θ2, α2, vB. The operational equation is linear in the parameters θ1, θ2, vB, and nonlinear in α2 . The θ1 and θ2 parameters are a combination of the rate constants.
Basis function method according to Hong and Fryer [1] in the irreversible case:
1.For a certain tracer the physiological range of the rate constants can be determined. These ranges are translated into a range of α2 values which can be expected in the data. With FDG, for instance, α2∈[0.06,0.6]min-1 was proposed in [1]. The range [0.06,2.4] is used as a default.
2.The basis functions ![]() are pre-calculated for tabulated α2 values which span the prescribed range.
are pre-calculated for tabulated α2 values which span the prescribed range.
3.When fitting data, each value of α2 is examined: the operational equation using the corresponding basis function is fitted with respect to the remaining parameters θ1, θ2, vB using the singular value decomposition method without weighting. Since all of them enter linearly, the solution is unique and can be quickly calculated. For each of the calculations the chi-square criterion is recorded.
4.Finally the combination θ1, θ2, vB, α2 with minimal chi square is considered as the solution.
It is possible to allow fitting of the blood volume fraction vB, or to fix it at a specific value. The configuration whether vB is fitted or fixed is specified in the preprocessing setup.
When setting up the processing it is recommended to enable calculation of the α2 map and inspect it regarding the prescribed range. If the prescribed maximum or minimum value is very frequently encountered this indicates that the range should be expanded.
Acquisition and Data Requirements
Image Data |
A dynamic FDG PET data set representing the measurements covering a sufficient time range after injecting of a 18F-Deoxy-Glucose (FDG) bolus. If the scanner FOV was not at a fixed location during the scan, the slice-times need to be contained in the image attributes. The model has been tested with the data of Siemens equipment (DICOM Conformance statement, see Frame Reference Time) capable of continuous table movement. |
Blood Data |
Blood activity from the time of injection until the end of the scan. It is important, that the blood and the image data have the same time base and decay correction is relative to the same time. |
Blood Preprocessing
The only necessary configuration is specification of the blood activity curve, either as a file, or as a VOI placed over a vessel such as the descending aorta or the left ventricle. Note, however, that the blood information must be available from the time of injection, whereas later measurements are sufficient for the tissue.
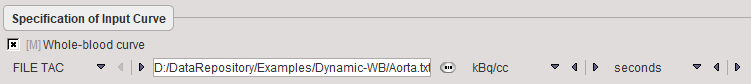
Model Preprocessing
The model processing panel includes two classes of options.
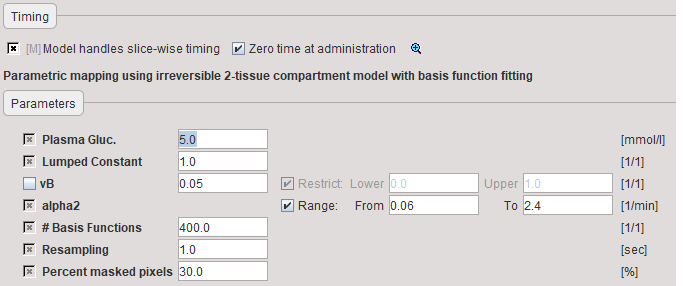
The options at the top are related to the handling of the timing, see Variable Timing of Slices. The actual model parameters are listed in the lower part.
Plasma Gluc. |
Plasma glucose in [mmol/l] measured with a blood sample of the patient. |
Lumped Constant |
The Lumped Constant is used to compensate for the difference in uptake between normal glucose and Fluoro-Deoxyglucose (FDG). It differs between tissues and therefore is set to 1 by default. |
vB |
Blood volume fraction. Can be fitted or fixed. If checked, the blood fraction is fitted during map calculation. Otherwise, the specified value will be used for spillover correction. |
alpha2 |
Second eigenvalue. For defining its basis function range first enable the alpha2 parameter and then adjust the Lower and Upper values. |
#Basis Functions |
Number of intermediate α2 values generated between Lower and Upper. The increments are logarithmically spaced. |
Resampling |
Sampling increment applied during the basis function calculation. |
Percent masked pixels |
Exclude the specified percentage of pixels based on histogram analysis of integrated signal energy. Not applied in the presence of a defined mask. |
Model Configuration
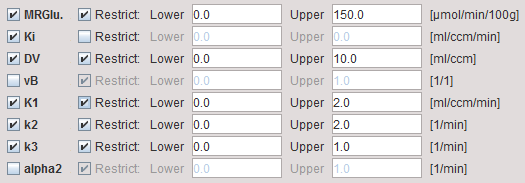
MRGlu |
Metabolic Rate of Glucose in [μmol/min/100ml], the actual result of the model. It is calculated as: |
Ki |
Influx Ki=(K1*k3)/(k2+k3) of the irreversible 2-tissue compartment model; is directly proportional to MRGlu. |
DV |
Distribution volume of FDG in tissue. |
vB |
Blood volume fraction defining the pixelwise blood spillover correction. To fit, please activate in the Preprocessing tab. Note, however, that the noise will be increased. |
K1,k2,k3 |
Rate constants of the irreversible 2-tissue compartment model. |
alpha2 |
Shows the α2 values corresponding to the basis function of the solution. Can be used to check whether the defined range was adequate. |
Reference
1.Hong YT, Fryer TD: Kinetic modelling using basis functions derived from two-tissue compartmental models with a plasma input function: general principle and application to [18F]fluorodeoxyglucose positron emission tomography. Neuroimage 2010, 51(1):164-172. DOI