For many of the supported models some type of preprocessing or the specification of some information piece is required before the actual pixelwise calculation can be started. In the Model Preprocessing panel, the preprocessing parameters of the current model are listed. Typically, there are input parameters which will also be used in the pixelwise calculation, as well as parameters of interest calculated during preprocessing.
Preprocessing operations may require certain input information, typically:
▪Tissue time activity curves: They serve for the calculation of initial parameters such as t*, and for checking that the model is working properly with the current data.
▪Fitable model parameters: Such parameters can potentially be fitted with the current data by enabling the fit checkbox ![]() and specifying the required time-activity curves. Otherwise, an already known value has to be entered and the fit checkbox disabled
and specifying the required time-activity curves. Otherwise, an already known value has to be entered and the fit checkbox disabled ![]() .
.
▪Input parameters ![]() : Here user input is requested, e.g. a Threshold percentage.
: Here user input is requested, e.g. a Threshold percentage.
The parameters of each model are described in a separate section of this document. Note the Set Defaults button which restores the configuration typically used with the model.
Logan Plot Example
In the case of the Vt (3 Calc Methods) model including the Logan plot illustrated below the equilibration time t* must be defined from which on the plot is considered linear and a regression line is fitted.
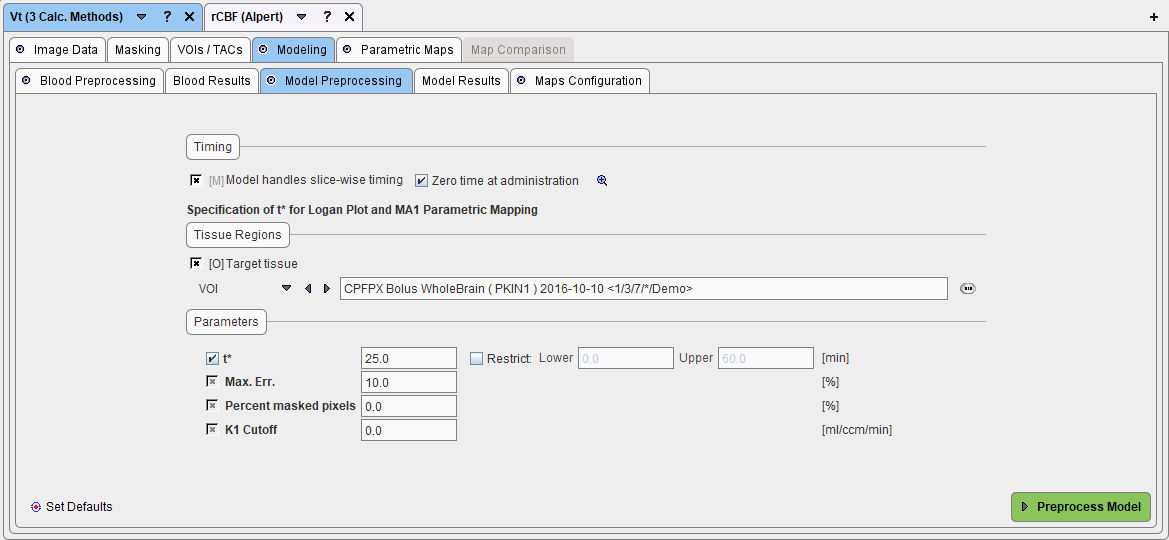
To determine t* with the current data the user has to do the following:
1.Enable the optional Target tissue checkbox and specify a tissue time-activity curve to which the Logan plot will be applied. This can be done by referencing a previously defined VOI as in the example above (VOI selection), by referencing a TAC data file with the FILE TAC selection, the DB TAC selection, or a THRESHOLD selection.
2.Enable the fit checkbox of t* in order to fit it using the error criterion specified in Max. Err.
The Percent masked pixels is a common input parameter in preprocessing which serves for background masking. The integrated signal energy is used as criterion, and the percentage defines the percentage of pixels in excluded based on a histogram analysis. A percentage of 30% means that the 70% pixels in the upper range of the histogram are retained. All excluded pixels will be masked to NaN in the parametric maps.
How To Continue
If all required information for the selected model has been specified, start the model preprocessing step with the Preprocess Model button.