Correction and calibration of twilite raw data is performed using the PSAMPLE TwiliteCorrection module from the PMOD ToolBox.
Please select the Correction page
and proceed as follows:
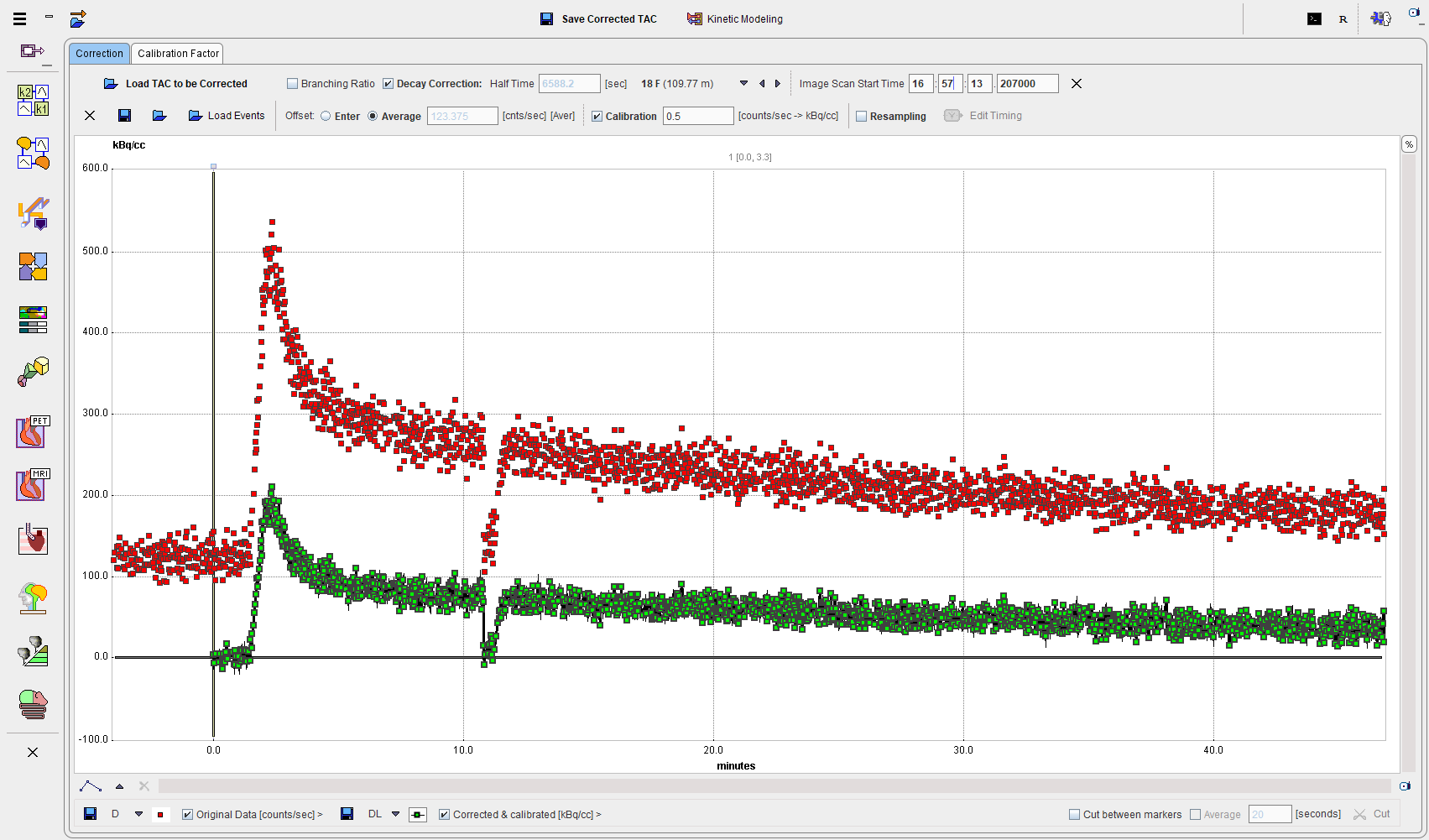
1.Load the raw data of the twilite experiment with the Load TAC to be Corrected button. The coincidence rate during the measurement is displayed in the curve area as Original Data [counts/sec] with red squares. It should show a level background at the beginning before the activity was injected, followed by a peak when the activity in the blood reaches the detector head.
2.Enable the Branching Ratio and Decay correction and select the appropriate isotope from the list.
3.Correct the PET Scan Start Time. Initially it is set to the time of the first twilite sample. Enter the proper PET start time corresponding to the start of dynamic imaging. Note that time 0 in the curve display is shifted to correspond to the difference between the start of twilite sampling and the true PET start time (i.e. the twilite is typically started before the PET acquisition, providing background count rate data). The decay correction will be applied relative to this time 0 to match the usual decay correction applied to PET data.
4.Select the Average option for background subtraction.
![]()
With this setting, all samples before time 0 are averaged, with the assumption that this is background signal only. The background average is shown in gray text and is subtracted from the Original Data before decay correction. If the twilite was not started before the PET acquisition, or the background was not stable before time 0 (e.g. high activity near the detector/light guides, such as 15O-water generation, Enter should be selected instead. The background level (e.g. that calculated during twilite calibration) should then be entered manually.
5.Activate the Calibration factor. Typically this factor has been established during the last calibration procedure, but can manually be overwritten. The calibration factor is multiplied with the background subtracted, decay and branching ratio corrected data, resulting in the Corrected & calibrated [kBq/cc] whole blood activity concentration. Note that the curve is truncated before time 0, assuming that data prior to PET acquisition is not required.
6.Save the calibrated whole blood activity concentration measurement using Save Corrected TAC. There are two formats available, from which the user has to choose in a dialog window.
![]()
The Blood format is used when the data will be loaded as a whole blood input curve into the PKIN tool (the typical use) and has only one time column, the sample mid-time. The Tissue format is intended for import of the data as a tissue TAC into PKIN, and therefore has a sample start and end time column.
7.As an alternative, the Corrected & calibrated curve can directly be transferred to PKIN via the Kinetic Modeling button. An interface window
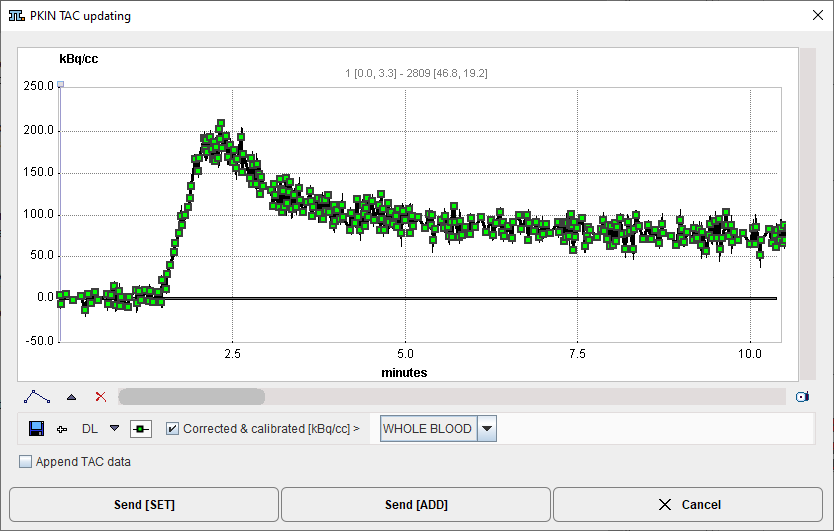
is opened which shows the curve and a drop-down menu to select how PKIN should interpret the data: as WHOLE BLOOD, PLASMA or TAC. Usually, WHOLE BLOOD should be selected. Send [SET] will overwrite data of the selected workspace in PKIN, whereas Send [ADD], will add a new PKIN workspace for the transferred data.