The template-based normalization for brain CT is an implementation of the SPM5 methodology. It adjusts the human CT input image to a human CT template image by applying an affine transformation first, followed by iterative elastic adjustments.
The Template-based normalization for BrainCT button provides access to the different types of elastic matching methods. An interface dialog window appears with two tabs. The first one serves to quickly retrieve species sensitive pre-defined or saved configurations of the iterative matching settings, the second one gives access to all parameters.
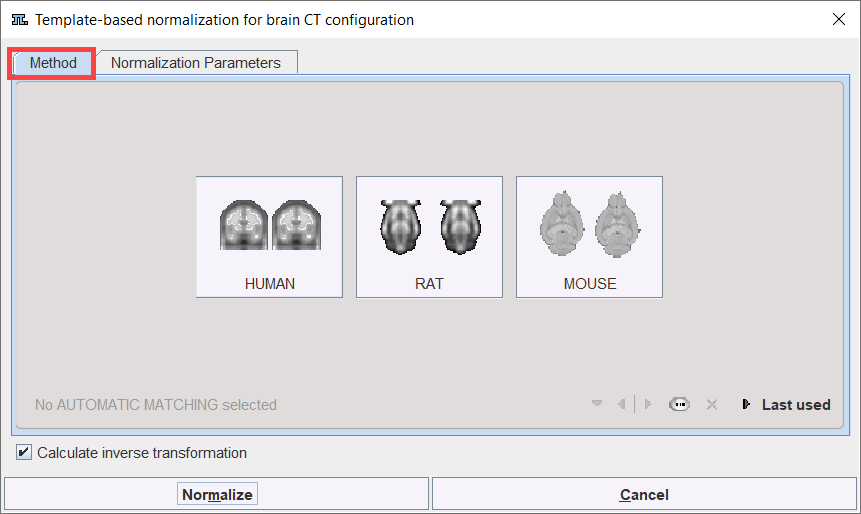
The human CT and CTCU templates are to be used with the Template-based normalization for brain CT matching method. These human brain templates provided with PMOD are all head first supine (HFS) oriented and are located at D:/Pmod4.3/resources/templates/normalization/
The following parameters allow fine-tuning the algorithm.
CT |
CT template for an older population created from 30 healthy individuals with ages similar to what is commonly seen in stroke (mean 65 years). Developed for the SPM8 Clinical Toolbox by Rorden et al [10]. |
CT CU |
CT template as above but with the converted units (CU) which improves the contrast for soft tissue and CSF so that the normalization procedure works better. Conversion procedure : HU values -1000..-100 are mapped to 0..900, values from -99..100 are linearly scaled to the range 911...3100, and values i>100 become [i+3000] [10]. CT images to be normalized with this template must also be converted by enabling the Transform HU values option. |
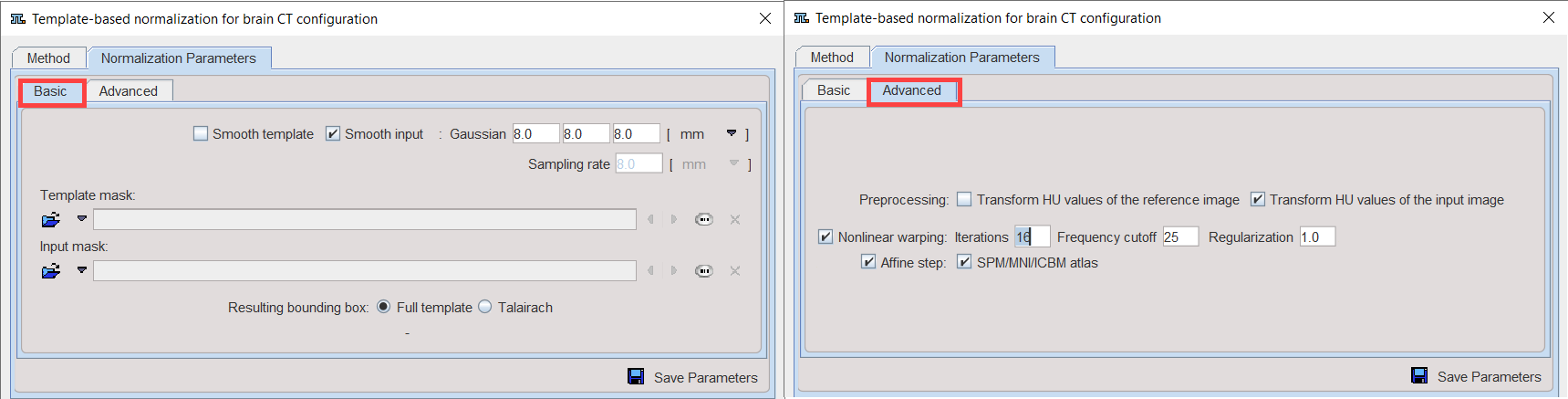
Note the Transform HU Values of [...] options which transform the values in the CT image such that the contrast between bone and soft tissue is reduced and they are more similar to the usual anatomical images.
Basic Parameters
Smooth template, |
If either box is checked, an initial Gaussian smoothing of the respective data is performed. Both smoothing operations use the same configurable FWHM parameters. Usually, the template has already been smoothed beforehand so its smoothing is normally not required for the normalization. |
Sampling rate |
The sampling rate of the method is derived from the Smooth Input filter size. If no smoothing is applied, the sampling rate needs to be specified by the user. |
Template Mask |
This selection allows defining a mask to be applied to the template during the normalization procedure. If one of the standard templates is used, its mask is implicitly defined and the selection is therefore inactive. |
Input Mask |
A mask file can be selected which masks the part of the input image which should be disregarded in the normalization. To discard a selected mask activate the Clear file button |
Resulting bounding box |
The radio box for defining the extent (bounding box) of the resulting normalized images. ▪Full atlas: The result image has the size of the template. ▪Talairach: The result image is trimmed to the bounding box of the Talairach brain atlas as in the SPM99 program. It is only applicable for MNI brain templates. |
Advanced Parameters
The Advanced parameters are usually only changed if a normalization fails or if the user aims at a specific effect.
Nonlinear |
If this box is not checked, only the affine (translation, rotation, scaling, shearing) part of the normalization is performed. |
Iterations |
Number of nonlinear iterations. The higher the iterations number, the more deformations may occur. |
Frequency cutoff |
The specified Frequency cutoff (default = 25) is used together with the Bounding box size to calculate the number of basis functions. Higher cutoff values result in fewer basis functions. |
Affine step |
Estimate and apply an affine transformation before the nonlinear warping iterations start. |
SPM/MNI/ICBM atlas |
Use settings which are appropriate for the templates of these standard atlases. |