Assumptions and Recommendations
PMOD includes a powerful database for the storage and organization of all kinds of relevant data. It is highly recommended that the data for Brain VOIs batch processing is organized in such a database. This will allow the user to take advantage of association between data elements (e.g. identification of PET/MR paired series) which can only be efficiently provided by a database. Furthermore, the output produced during batch processing can directly be inserted at the appropriate database level, which is more difficult when working with directory structures.
Association of MR and PET Series
The PMOD database alows the user to define a relationship between images, and other components, called "association". For the analysis of hybrid data such as PET and MRI it is convenient to associate the two image series, because thereafter it is sufficient to select only one of the images in a workflow that will utilise both.
Association of image pairs can be performed on the DB Load page of PNEURO as illustrated below.
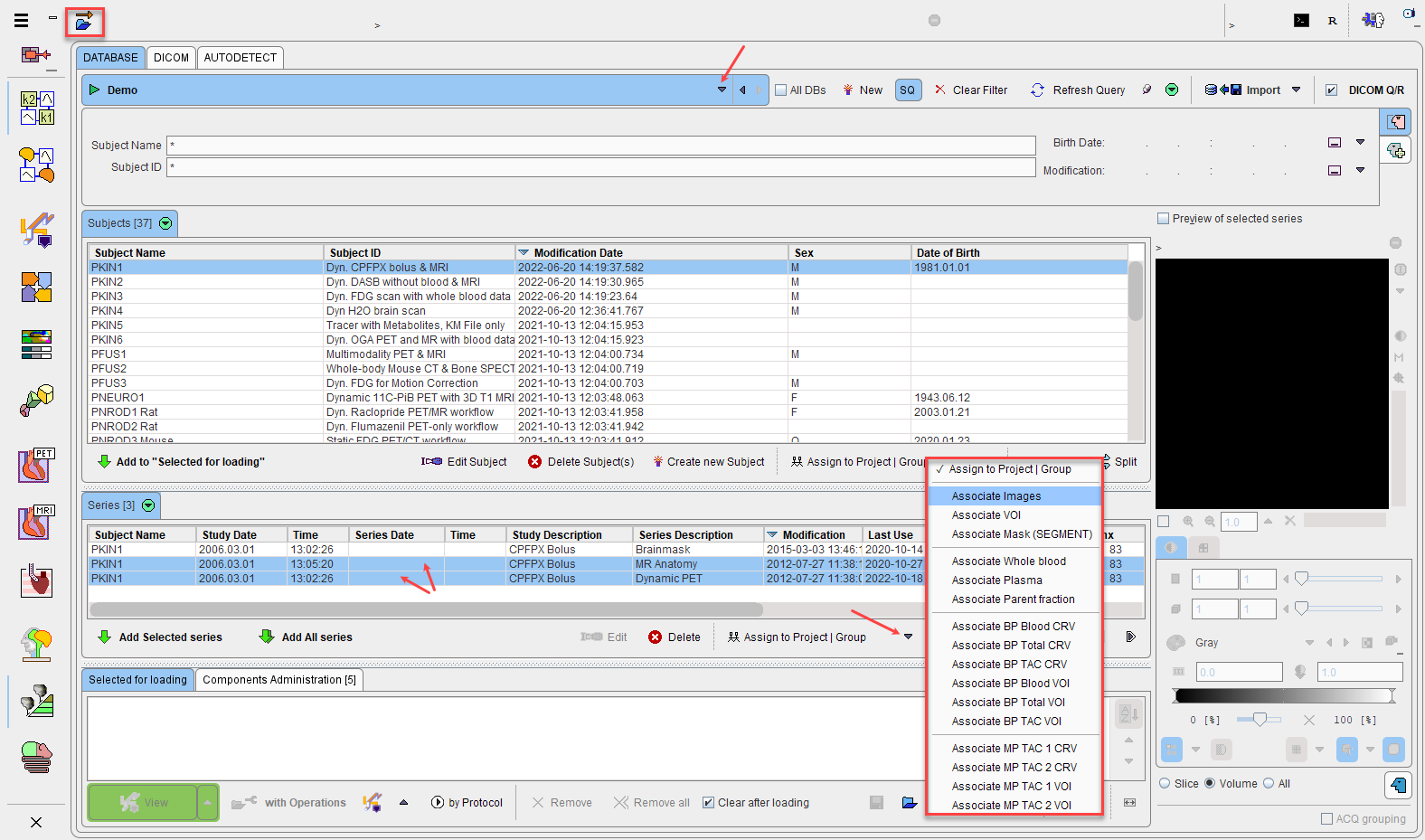
Select a subject in the Subjects list. In its Series list select the two image series to be associated. Finally open the Set Project option menu and activate the Associate Images entry to establish the mutual relation between the image series. This operation has to be repeated for all subjects to be processed. It is a one-time action which however can be taken advantage of in multiple analysis procedures.
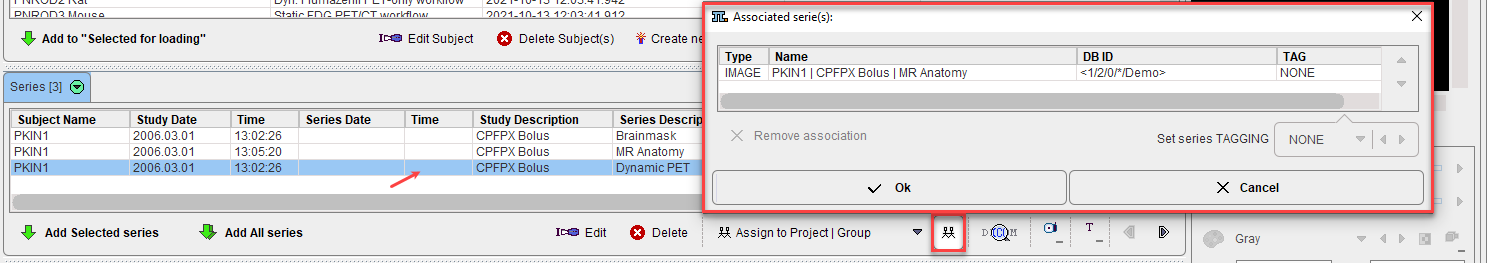
The image(s) associated with a given series can be reviewed using the List Associations button:
Batch Process Overview
The basic concept of the batch mode is the sequential processing of saved protocol files. Such protocols can be created by loading the data sets (PET and/or MR), defining crop boxes and crop the images, adjusting the processing parameters on the different pages, and saving a protocol file. An alternative is to prepare a master protocol file which can be applied for generating individual protocol files for a whole set of (associated) input data. This procedure is described below.
Starting the Batch Brain VOIs Procedure
Use the menu entry Batch Mode/Maximum Probability Atlas & Brain Parcellation (Brain VOIs)
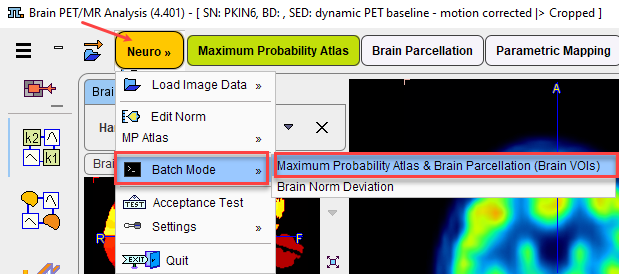
or the button ![]() from the lateral task bar to start the batch operation.
from the lateral task bar to start the batch operation.
First, a dialog window appears for selecting existing PNEURO protocol(s) (.pbrainProt) from a database. Select the appropriate protocols and activate Set selected. Use Set from File System to load protocols from outside the database.
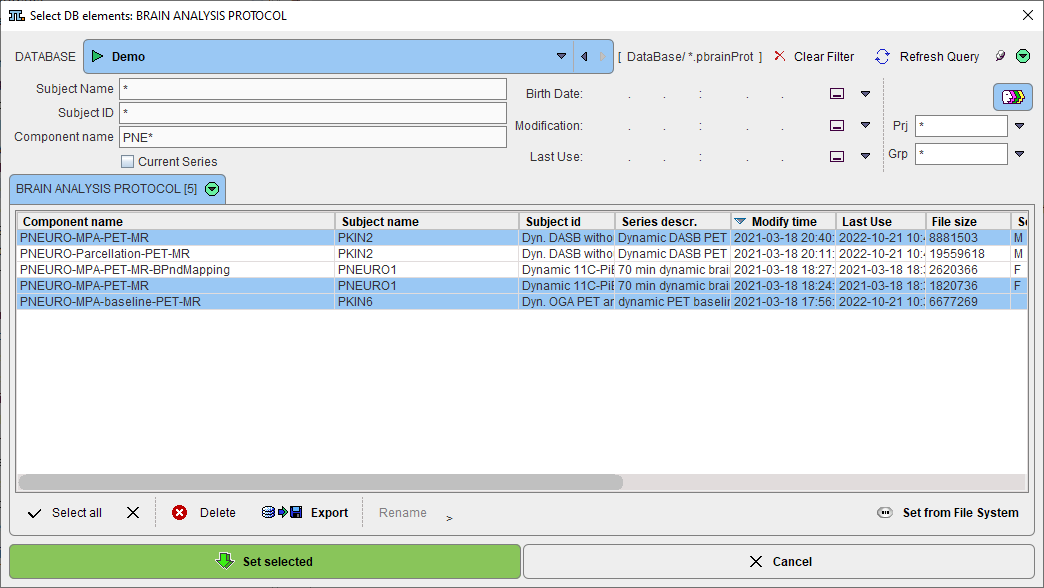
Note that the method for the VOI generation is defined from the protocol and can't be changed, only its parameters
The Batch Brain VOIs Interface:
The interface of the Batch Brain VOIs is illustrated below:
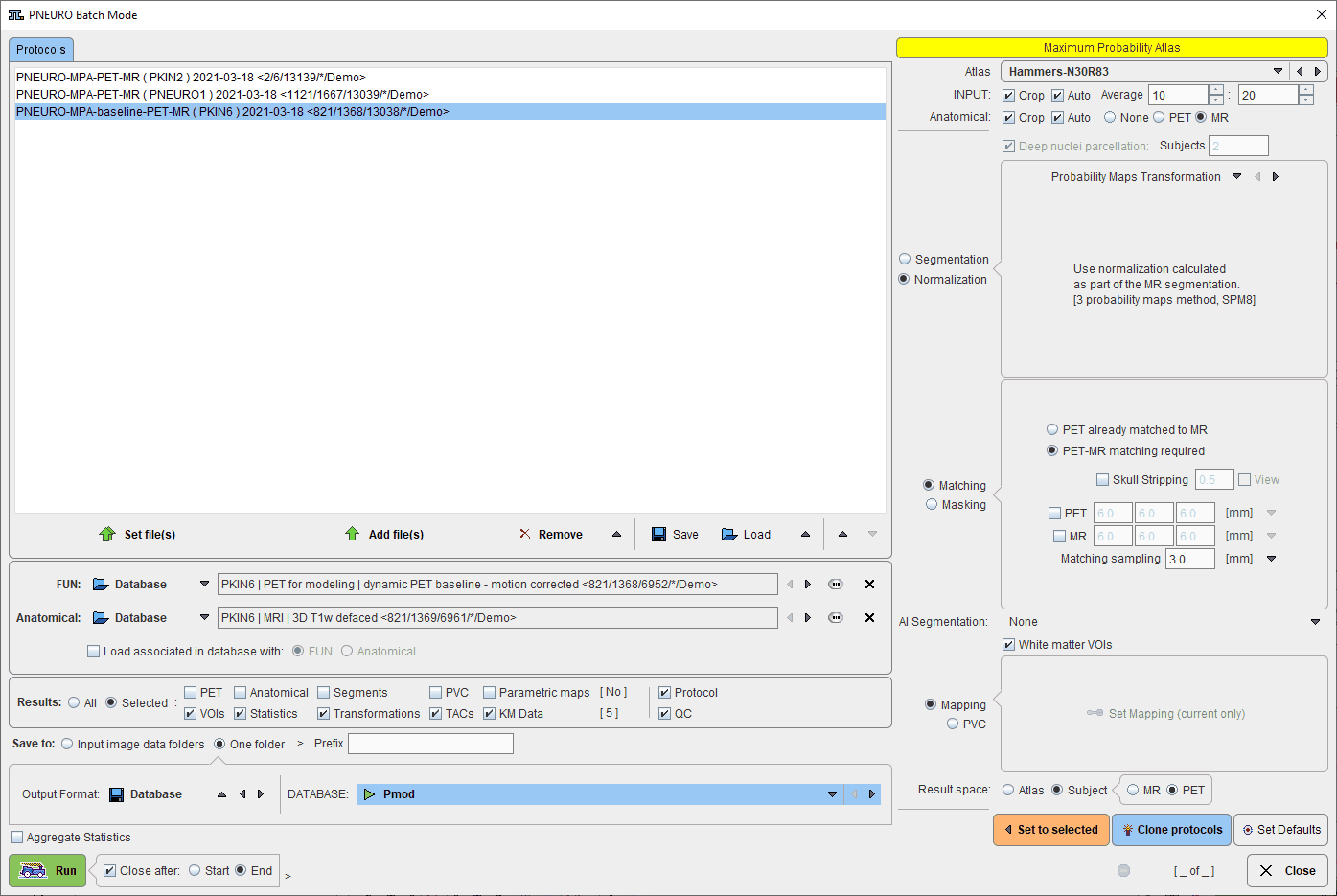
The window has four areas, which are described below.
Protocols List
The list of the protocols which will be processed is displayed in the upper left. Additional protocols can be added with the Add file(s) button, and selected entries removed from the list by Remove. When an entry is selected in the protocols list, its data definition is updated in the FUN and Anatomical text field, and its processing parameters are shown to the right.
Parameter Settings
The parameters of the protocol (or an entire set of selected protocols) can be edited in the area to the right and then updated with Set to selected protocols. This mechanism makes it easy to establish a common processing algorithm.
Note: The Deep nuclei parcellation and Subjects definitions are active only when a brain parcellation protocol is selected on the protocols list.
Cloning Protocols
Once a protocol with suitable processing parameters has been defined, it can be cloned for application with additional data sets. This is achieved by selecting the template protocol in the list and activating Clone protocols the lower right area. A dialog window opens for definition of the input data.
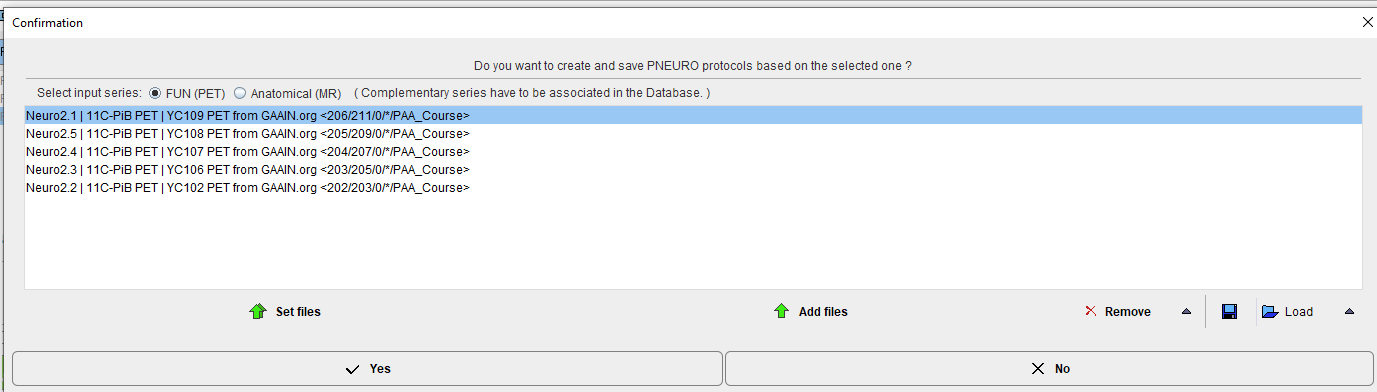
Depending on the type of analysis, the corresponding radio button FUN (PET) or Anatomical (MR) has to be configured. For workflows with both MR and PET images, the image series have to be associated beforehand as described above.
Please select the appropriate image files with Set files, and append additional ones with Add files. Remove deletes a selected entries from the list, whereas Remove all clears the whole list. An input data list can be saved for later use with the Save button, and retrieved with Load.
Once the data list is complete, activate Yes to generate a new protocol list containing a protocol for each of the input data sets. The protocols are saved in the database and associated with the corresponding images.
Note: If cropping is used in cloning, it is recommended using the automatic placement of the crop box by enabling the Crop and Auto boxes in the parameter settings area.
Results and Output Configuration
The granularity of the results saved during the batch processing can be configured in the Results section. With All enabled, all possible results are stored at the end of processing. With Selected enabled, the following components can be enabled for saving:
▪PET (in final space): PET image in the space where the VOIs are generated.
▪Anatomical (in final space): The anatomical image (MR or FDG) in the space where the VOIs are generated.
▪Segments: The result of the tissue segmentation into grey matter, white matter, CSF.
▪PVC: The image results of the partial volume correction configured in the VOIs page.
▪Parametric maps: The parametric maps results obtained from the mapping procedure. Please note that only the protocols saved at the end of the mapping procedure will generate parametric maps.
▪VOIs: Contour VOIs in the selected result space.
▪Statistics: VOIs applied to the target image series (PET, if available, otherwise the anatomical image).
▪Transformations: The transformations calculated based on the protocol specific workflow .
▪TACs: The TACs statistics in the selected result space.
▪KM Data: The TACs are saved as a .kmData file ready to be used in the PKIN processing.
▪Protocol: The protocol after the processing is (re)saved in the specified loaction. This is relevant for the situation when the list of protocols was created using the cloning facility. Such protocol includes definition of the input data as well as the parameters of the different processing stages, including the parametric mapping settings when available.
▪QC: The quality control capture is generated.
The output destination is set with the Save to configuration. A meaningful Prefix for the saved data components is recommended. The destination configuration is only relevant when saving to the file system. In that case, the results can be distributed to the Input image data folders, or concentrated in One folder, which needs to be specified together with the output image format.
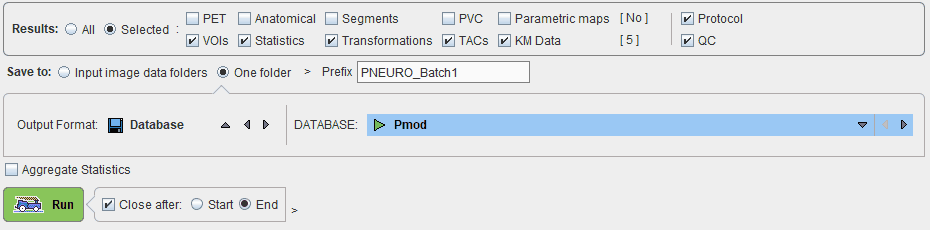
Note: The time activity curves generated during parametric mapping protocols may also be saved by enabling the TACs check box or by enabling the save KM Data check box.
The Aggregate Statistics option is relevant when the batch is setup to run protocols for the same group and condition. If the option is enabled, the results of all data sets are concentrated into a single aggregation table, which can immediately be used for statistical analysis.
Note: In case of multiple parametric mapping statistics, only one type of statistics should be selected for aggregation at the end of the processing.
Run the Batch
The batch processing is started with the Run button.
![]()
To run processing without the batch window open (recommended) configure Close after Start. While processing is performed, PNEURO will remain blocked and run through the processing steps for each protocol. The Aggregate Statistics option is convenient for assembly of the resulting VOI statistics output into an aggregate file for subsequent statistical analysis. In case of multiple parametric mapping statistics files only one type should be selected for the direct aggregation form batch.
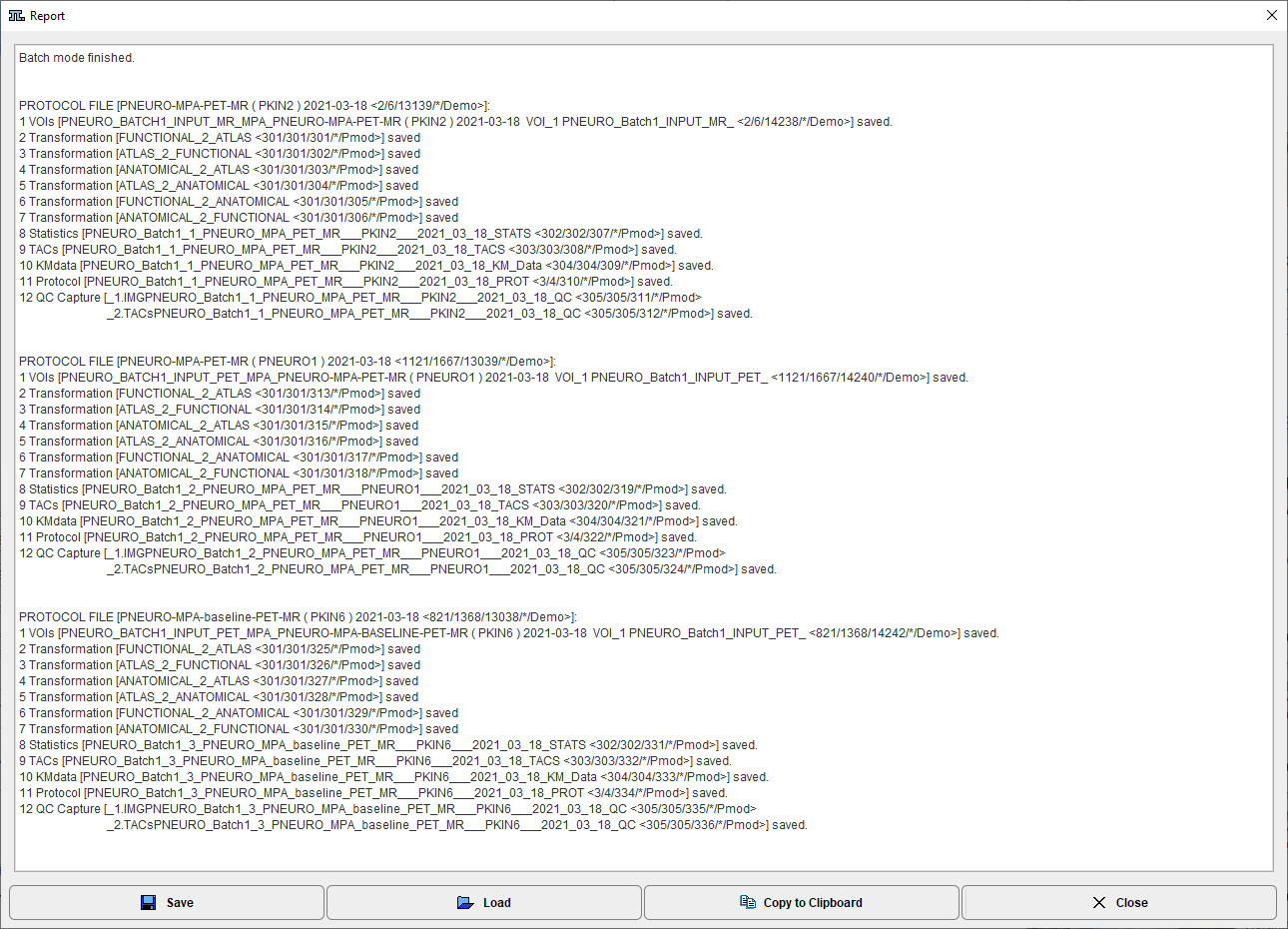
After batch processing has completed, a summary is shown in a Report window detailing the saved results as well as potential errors. The report can saved using the Save button, and added t the clipboard using the Copy to Clipboard button.